Columbia University
Irving Medical Center
Neurological Institute
710 West 168th Street, 3rd floor
(212) 305-1818
TaubCONNECT Research Perspectives:
January 2017
#2 Novel Genetic Loci Underlying Human Intracranial Volume Identified Through Genome-Wide Association
#3 Relation of Dysglycemia to Structural Brain Changes in a Multiethnic Elderly Cohort
 |  | |
| S. Abid Hussaini, PhD | Karen Duff, PhD |
|
 |  | |
| Hongjun (Harry) Fu, PhD | Gustavo Rodriguez, PhD |
One of the earliest cognitive symptoms of Alzheimer's disease (AD) is confusion and disorientation in familiar surroundings. The entorhinal cortex (EC) is one of the regions in the brain known to store information of the space around us to create a 'cognitive map' and neurons of EC are among the first to be affected in AD. To understand how AD pathology leads to impaired spatial memory and cognitive map dysfunction, Drs. Karen Duff, Abid Hussaini, Hongjun (Harry) Fu, Gustavo Rodriguez, and colleagues from the Duff and Hussaini laboratories studied a mouse model (EC-Tau) which replicates the earliest stages of human Alzheimer's disease by forming Tau pathology in the EC that then spreads to the hippocampus and neocortex as the mice age.
Published this month in Neuron, the team found that the old (30+ month) EC-tau mice had significant difficulties navigating the Morris water maze and the T-maze compared to younger EC-tau mice with less pathology and controls. To investigate which neurons were involved in the observed spatial memory impairment, Drs. Hussaini and Rodriguez used in vivo electrophysiology to record neuronal activity in the EC as the mice explored an open field box. Extracellular electrophysiology recordings from the EC allowed them to evaluate the properties of excitatory as well
as inhibitory neurons.
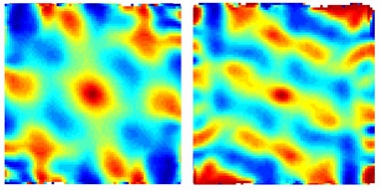
Figure: GPS Fail. The grid cell firing pattern in a mouse with tau pathology in the entorhinal cortex (right) is weaker than in an age-matched control (left). [Courtesy of Abid Hussaini, Columbia University.]
Among the excitatory neurons, a specialized group of cells, called the grid cells, were affected. These recently discovered cells are like a coordinate or a map of the brain's GPS system and are known to be important for spatial memory. Normally, the grid cells fire uniformly across the entire space of the environment in a grid-like periodic pattern, but in the old EC-tau mice these cells fired less and the grid patterns were less periodic. Other excitatory cell types in the EC, including head-direction and border cells, were not affected. The inhibitory neurons had an opposite effect, firing a lot more in the old EC-tau mice than in the control group. These opposite effects changed the excitatory-inhibitory balance in the neuronal population of EC, most likely altering the overall network activity. This imbalance led to an increase in theta wave frequency which is known to be increased in human AD.
Detailed histopathological studies led by Dr. Harry Fu showed that tau pathology in the EC only formed in excitatory cells but not inhibitory cells, which were completely spared. Furthermore, overt tau pathology in the older mice was linked to extensive cell death, which only depleted the excitatory neurons but not inhibitory neurons.
These data suggest that deficits in spatial memory seen in early Alzheimer's disease can be caused by disruption of the brain's GPS system due to the selective vulnerability of excitatory neurons in the EC region of the brain. Tests of spatial memory, such as those being developed for smart-phone use by the elderly, might reveal impairment earlier than cognitive tests that are currently used. Results from this study by Fu et al. were also featured on ALZFORUM and CUMC Newsroom this month.
S. Abid Hussaini, PhD
Assistant Professor of Neurobiology (in Pathology and Cell Biology and the Taub Institute)
sah2149@cumc.columbia.edu
Karen Duff, PhD
Deputy Director, Taub Institute
Professor of Pathology and Cell Biology (in Psychiatry and the Taub Institute)
ked2115@cumc.columbia.edu
Hongjun (Harry) Fu, PhD
Associate Research Scientist (in the Taub Institute)
hf2296@columbia.edu
Gustavo Rodriguez, PhD
Postdoctoral Research Scientist (in the Taub Institute)
gr2501@cumc.columbia.edu
Novel Genetic Loci Underlying Human Intracranial Volume Identified Through Genome-Wide Association
 |  |  | ||
| Sandra Barral, PhD | Adam M. Brickman, PhD | Badri N. Vardarajan, PhD, MS |
Intracranial volume reflects the maximally attained brain size during development. It is closely related to brain volume in early life as the brain grows, but becomes stable after the brain has fully developed and remains unaffected by later age-related changes such as brain atrophy. It is a highly heritable trait, complemented by environmental factors, and identifying the underlying genes that influence variation in intracranial volume can contribute to our understanding of human brain development and related diseases. Yet, these genes have remained challenging to discover. Prior studies have identified only two influential genetic loci.
Drs. Sandra Barral, Adam Brickman, and Badri Vardarajan from the Taub Institute recently participated in a worldwide project of unprecedented scale, in which the largest-ever meta-analysis of a genome-wide association study (GWAS) of intracranial volume of 32,438 adults was performed. As published in Nature Neuroscience this month, five previously unknown genetic loci associated with intracranial volume were discovered and two known signals confirmed. According to the authors, the genes in these loci provide intriguing links between maximal brain size and various processes, including neural stem cell proliferation (FOXO3), neurodegeneration (MAPT), bone mineralization (CENPW), growth signaling (IGF1, HMGA2), DNA replication (GMNC) and rRNA maturation (PDCD). On a genome-wide scale, they discovered evidence of genetic correlation between intracranial volume and other key traits such as height and cognitive function, as well as with Parkinson's disease, indicating that the genes underlying brain development have far-reaching effects that extend well beyond the initial years of life. Their findings identify the biological underpinnings of intracranial volume and their link to physiological and pathological traits.
For this study, Drs. Barral, Brickman, and Vardarajan directed the genetic and neuroimaging analysis of the Washington Heights-Inwood Community Aging Project (WHICAP) cohort, a community-based longitudinal study of aging and dementia among elderly, urban-dwelling residents led by Taub Co-Director Dr. Richard Mayeux since 1989. Drs. Barral and Vardarajan conducted genetic analysis to identify variants that influence intracranial volume using the genome-wide imputed genotyped data available. Their research integrates different statistical approaches (linkage analysis, case-control analysis, genome-wide association analysis, and whole exome/genome sequencing) to comprehensively characterize associations between genetic variants and neurological phenotypes, in studies of both normal aging and diseases of aging, such as Alzheimer’s disease. Dr. Brickman has been directing the acquisition and analysis of high-resolution magnetic resonance imaging (MRI) data from WHICAP. His laboratory primarily focuses on quantifying markers of neurodegeneration, normal brain integrity, and cerebrovascular disease, including measures of intracranial volume.
Sandra Barral, PhD
Assistant Professor of Neurogenetics (in Neurology, the Taub Institute, and Sergievsky Center)
smb2174@columbia.edu
Adam M. Brickman, PhD
Associate Professor of Neuropsychology (in Neurology, the Taub Institute, and Sergievsky Center)
amb2139@cumc.columbia.edu
Badri N. Vardarajan, PhD, MS
Assistant Professor of Neurological Science (in Neurology, the Gertrude H. Sergievsky Center, and the Taub Institute)
bnv2103@cumc.columbia.edu
Relation of Dysglycemia to Structural Brain Changes in a Multiethnic Elderly Cohort
 |  |  | ||
| Christiane Reitz, MD, PhD | Adam M. Brickman, PhD | Jose Luchsinger, MD |
Dysglycemia is one of the most common public health problems in the United States. According to 2011 prevalence data from the Centers for Disease Control and Prevention (CDC), diabetes affects 25.8 million people in the United States, corresponding to 8.3% of the total U.S. population. Seventy-nine million persons (i.e. more than a quarter of the U.S. population) have pre-diabetes. This problem is more common in the elderly, the group also at highest risk for cognitive impairment. While there is evidence from epidemiological studies suggesting that dysglycemia is a risk factor for cognitive impairment, the underlying pathophysiological mechanisms remain unclear.
Published online in the Journal of the American Geriatrics Society, Drs. Christiane Reitz, Adam Brickman, Jose Luchsinger, and colleagues examined the relation of dysglycemia to brain structure and cognitive function in older adults in the Washington Heights-Inwood Community Aging Project (WHICAP) cohort. In this study, dysglycemia (diabetes, undiagnosed diabetes, and pre-diabetes) or higher HBA1c levels was associated with a higher number of brain infarcts, WMH volume and decreased total white matter, gray matter volumes and hippocampus volume in cross-sectional analyses, and a significant decline in gray matter volume in longitudinal analyses. In addition, dysglycemia was associated with lower performance in language, speed and visuospatial function, although these associations were attenuated when adjusting for education, APOE genotype, ethnic group, or vascular risk factors. These findings confirm that dysglycemia and diabetes affect cognition through both vascular and neurodegenerative pathways. Drs. Reitz, Brickman, and Luchsinger are conducting studies in humans and mice to further examine the mechanisms for these pathways.
Christiane Reitz, MD, PhD
Assistant Professor of Neurology and Epidemiology (in the Taub Institute and Sergievsky Center)
cr2101@columbia.edu
Adam M. Brickman, PhD
Associate Professor of Neuropsychology (in Neurology, the Taub Institute, and Sergievsky Center)
amb2139@cumc.columbia.edu
Jose Luchsinger, MD
Associate Professor of Medicine and Epidemiology (in the Taub Institute)
jal94@columbia.edu